Steven Benner, a synthetic biologist, and his team have synthesized four new letters of DNA. The study was published in the journal of Science. The new eight letter genetic system is being called "hachimoji." Researchers used the four natural letters of DNA as a model, then used chemistry to design and synthesize four new letters that are the same size and shape as the originals. According to the researchers, the new genetic system meets four of the five requirements for life. The four they meet are; information storage, information transfer, selection, and evolvability. The researchers also synthesized a RNA molecule that transcribes their synthetic DNA. The fifth requirement that the system does not meet is self-sustainability. It does not reach this requirement because it does not exist in an organism, it exists only in a test tube in their lab that must be maintained by a graduate student. This also means it does not pose any harm outside of the laboratory.
I believe this discovery is very important in the field of genetics, as well as the field of medicine. The researchers have already began testing the new system to bind to cancer cells and anthrax toxins. With further studies and more time to work on the molecules, this new genetic system could bind to toxins in the body and destroy pathogens.
Showing posts with label synthetic biology. Show all posts
Showing posts with label synthetic biology. Show all posts
Tuesday, April 9, 2019
Wednesday, April 19, 2017
Simplified life
There has been recent attempts to create an organism with the smallest genome possible. Previously, the mycoplasmas, a genus of bacteria, were identified as forms of life with minimal genetic code. These bacteria, which lack a cell wall, have been proposed as ideal model organisms to study as a means of understanding the basic genetic requirements essential for life. Mycoplasma genitalium was sequenced and revealed the smallest genome of any autonomously replicating cell found naturally (525 genes). Recently, scientists have tried to create an organism with an even smaller genome with the rationale that all of Mycoplasma genitalium’s 525 genes were not essential and if some could be stripped away, it would lead researchers to discover the absolute minimum code required for life. After manipulation, a synthetic strain of Mycoplasma labeled (JCV-syn1.0) was created with only 473 genes. Although only ~250 genes appeared to be essential for proper growth, it was discovered that there may be quasi-essential genes that have implications in robust growth of this bacteria. There were also 149 genes that had unknown biological functions yet appeared important when creating this bacteria strain. This work is intriguing and will hopefully lead us to a better understanding of the building blocks of life. Although this is an attempt to simplify life, the results turned out to be very complex.
Article:
Pop news article:
Videos:
Tuesday, April 18, 2017
First Synthetic Genome
Researchers are coming very close to finishing the first fully synthetic genome. Nearly 3 years after the team revealed the first "designer chromosome" they now have 6 of the 16 chromosomes they need to complete the genome of baker's yeast. Theses chromosomes are not an exact copy but a cut-up and stitched together assembly of the coding portions of the known Saccharomyces cerevisiae genome. According to the Discover article the resulting genome "has roughly 1 million nucleotide-level differences from the natural version."
The project will hopefully yield a single functioning yeast cell containing a fully synthetic set of chromosomes. If this end result is achieved it will give us insight into aspects of the genome of Baker's yeast and even our own. We may be able to identify the role of non-coding regions since for the most part they are being removed in this project. If nothing else, as the author stated, this is an important step in the field of synthetic biology.
The project will hopefully yield a single functioning yeast cell containing a fully synthetic set of chromosomes. If this end result is achieved it will give us insight into aspects of the genome of Baker's yeast and even our own. We may be able to identify the role of non-coding regions since for the most part they are being removed in this project. If nothing else, as the author stated, this is an important step in the field of synthetic biology.
Friday, November 18, 2016
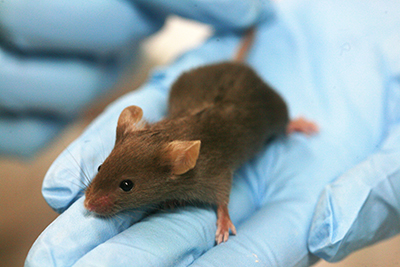
Genome editor CRISPR could put mutant mice in everyone's reach
Jackson Laboratory in Bar Harbor, Maine, genetically engineers model laboratory mice which it sells to researchers under the brand name JAX mice. Conventional development of new and modified mice strains for specific research has long relied on the laborious and multi-step process of small modifications to embryonic stem cells over the course of several generations. This process, which generally takes up to 2 years, is now being replaced by the CRISPR method of modification.
Using CRISPR, a targeted "genetic surgery" may be conducted on a fertilized egg. The length of time necessary to create a modified strain has been cut down to 6 months. These genetically engineered mice, modified to "knock out" or "knock in" genes are employed as key research models for diseases such as alzheimer's, osteoarthritis, muscular dystrophy, and Parkinson's.
Using CRISPR, a targeted "genetic surgery" may be conducted on a fertilized egg. The length of time necessary to create a modified strain has been cut down to 6 months. These genetically engineered mice, modified to "knock out" or "knock in" genes are employed as key research models for diseases such as alzheimer's, osteoarthritis, muscular dystrophy, and Parkinson's.
CRISPR has also significantly reduced the cost necessary to modify mice. Tak Mak, of the University of Toronto in Canada, who was also a pioneer in the mouse-mutating business, estimates it's about 30% cheaper to engineer a mouse with CRISPR instead of ES cells. At prices up to $20,000 USD for a custom mouse strain from JAX, this price reduction, as well as the remarkably easy technique, has the potential effect of opening up smaller labs to simply modifying strains themselves.
An international consortium consisting of a number of model mice development companies, including JAX, is currently working towards knocking out all 21,000 mouse genes, one by one, in order to reveal their function. The US National Institute of Health is so impressed by CRISPR's cost and ease of use, that it no longer funds consortium investigators to use embryonic stem cells.
CRISPR, as well as further developments in genome editing and synthetic biology, are my primary area of interest. I hope to find a career in which I may be able to work with and learn more about these systems.
Labels:
CRISPR-Cas9,
genome editing,
model organism,
National Institute of Health,
synthetic biology
Thursday, May 5, 2016
Artificial Antibodies and Antivenom
Another way geneticists and chemists have tried helping the costs go down is to develop synthetic antibodies. Using synthetics rather than waiting for animal plasma and their antibodies to take in the human host means faster administration and quicker healing. Synthetic versions of antibodies can be much smaller than the naturally occurring variety, which can help the victim by dropping the need for animal plasma and saving costs that would be used for maintaining and collecting animal plasma. In today's time there are a little over 90,000 deaths that occur each year because the treatment in so hard to come by people in both underdeveloped and developed countries. Africa today is one of the hardest hit by snakebite troubles. It has been reported that their supply of antivenom will likely be depleted sometime within this year. Snakebites don't get publicity as much as other tragic and traumatic events, but the threat to human life is real. And the effects of snakebites sometimes damage people for life, if they survived the bite at all that is. As we progress the techniques of synthetic biology I feel like we can find ourselves in a much healthier world. It's going to take time, but we have been making strides into more advanced techniques like we see here. The possibilities can be limitless.
Labels:
antivenom,
artificial antibodies,
DNA,
synthetic biology
Monday, April 11, 2016
Minimal Cell Created with just 473 Genes.
At the J. Craig Venter institute In La
Jolla, California scientist have created a “minimal cell” named
JCVI-syn3.0. This created cell, based on the Bacterium Mycoplasma
mycoides, has a synthetic genome with just 473 genes. While there was
worry that stripping down a cell to it's bare essentials would leave
it defenseless the JCVI-syn3.0 seems to be thriving in it's given
environment. The notion of this minimal cell was discussed at the
first international meeting on synthetic biology at Massachusetts.
They developed methods for chopping up
the bacterium's genome and removing segments piece by piece, allowing
them to get just the essentials of the cell. The starting point for
this minimal cell was an earlier synthetic cell, JCVI-syn1.0. With
techniques like this they hope to be able to redesign bacteria and
other relatively simple organisms so that they generate useful
chemicals and materials. For more information on the minimal cell
here and here.
Sunday, March 27, 2016
New Lightweight Champion of Genomes
Chris Venter, a genome sequencing pioneer of San Diego, California, and his team have released a report describing their recently engineered bacteria with the smallest genome of any freely living organism ever recorded.
This synthetic organism has been strapped down to only the bare minimum of potentially the only genes necessary to survive and reproduce. Unlike the genome of 20,000-25,000 possessed by humans, or the heavyweight champion Japanese flower that is comprised of 50 times more DNA than humans, this new bacteria known as Syn. 3.0 is comprised of only an astounding 473 genes.Venter's previous work had created the Syn 1.0, which had a genome of 901. In an attempt to shrink the genome smaller than the leading minimal genome of 525 in Mycoplasma genitalium, Vetner's large team of colleagues broke up into teams for each section of the bacteria's genome and kept stripping genes from Syn 1.0 with functions that were either nonessential or duplicated the function of another gene until they had a viable organism with a smaller genome than the previous leader.
Venter claims him and his team designed and tested "multiple hundreds" of constructs until they settled on Syn 3.0. Compared to M. genitalium which can take weeks for a population of its cells to double, Syn 3.0's slimmed down genome is able to double in 3 hours. The function of 149 of Syn 3.0's 473 genes still remain unknown, which leads way to further investigations and insights into the basic biology of life. Evolutionary biologists and biotechnologists are claiming to also start adding genes back to the genome of 3.0 to study their effects.
This is extremely exciting stuff, as it basically tells us that biologists have been paved a path for new discoveries. Syn 3.0 seems like the key to a new doorway of genetics which could potentially help lead to discoveries that just might end up in the school textbooks of the future.
Sunday, November 22, 2015
To Program Your House Plants
All the
computers and electronics we currently are just a complex arrangement of
simple, modular parts that control specific functions; well scientist at
Colorado State University are using the same modularity in plants by designing
gene that can control specific characteristics—color, size resistance to drought
or oxygen production. There is a
relatively new interdisciplinary field
is synthetic biology, the design of genetic circuits that control different
functions and can be easily placed in one organism or the next. Most of today’s
synthetic biologist only work with microorganisms like E. coli or yeast due to
the speed of reproduction and expression of gene but now they are moving onto
more complex organisms.
Using
plants comes with another problem for synthetic biologist, the complexity of
the organism they are instead altering 100 genes in the plant instead of one or
two like in single cell microorganism. They
are using protoplasts to alter the plant’s genes.
This
experiment could lead to a lot of different uses. First farmers could get grains and plants
that are more resistance to drought and by used in area where there are less
rainfall. NASA could develop plants can
be planted on another planet like Mars and live in the atmosphere unprotected
and then years later could produce oxygen on Mars to make it livable to humans.
I think the most used application of these techniques with me consumers to buy
plants and tree that could be programmed to randomly change colors or scents so
that it won’t clash with some rich person’s home décor.
Monday, April 20, 2015
Millions of Liters of Expensive Juice from One Fruit
Nootkatone is an expensive substance that costs more than $4,000 per kilo and can only be found as an aromatic in small quantities within grapefruits. Nootkatone is used in many different industries for a variety of different things. It can work as an insecticide, actively works against cancer cell lines in medicines, it has a nice smell for beauty products, and is even used in soft drinks for a subtle taste.
"We have installed new genetic information in the yeast Pichia pastoris, so that our cells are able to produce Nootkatone from sugar," says Austrian Centre of Industrial Biotechnology lab reasearcher Tamara Wrlessnegger. The yeast cells had their genomes altered with the addition of four foreign genes from the cress Arabidopsis thaliana, the Egyptian henbane Hyoscyamus muticus, the Nootka cypress Xanthocyparis nootkatensis and yeast Saccharomyces cerevisiae. The aroma from a single grapefruit is then used to create millions of liters of this functional juice.
I think the use of synthetic biology to solve the problem of acquiring this expensive substance is brilliant. Not only is the industrial and monetary value from an experiment like this great but it give these researchers a chance to perform a practical application of using cells to produce compounds for everyday use. This reminded me of a recent genetics video where scientists used E. coli to produce synthetic spider silk which is just another example of how synthetic biology is being used today. I hope that these types of experiments continue to provide the world with an even greater quality of life from the different substances that can be synthesized.
Spider Silk Video: http://www.sciencedaily.com/videos/4b4fe394e8729a35a97a00658bf145c9.htm
Sunday, November 17, 2013
Synthetic Creatures could “save nature” says Alexandra Daisy
Ginsberg

 In a futuristic proposal called Designing for the Sixth
Extinction explores the idea of putting synthetic living creatures into the
wild to save endangered species and clean up pollution. Ginsberg states, "The idea is that we could preserve or maintain a
state of nature using synthetic organisms that are designed to save other
species." Four new species have been proposed including a slug that
leaves a trail of alkali to neutralize acidic soil and a porcupine with sticky
rubber spines that would help disperse seeds of threatened plants. There is also an artificial puffball that
kills tree-damaging pathogens when it bursts; and a biofilm that grows on
leaves and absorbs pollutants and viruses, safely removing them when the leaves
fall in autumn. The creatures would be engineered to contain a genetic
"kill switch" that would prevent them from over-breeding and creating
new environmental problems.
In a futuristic proposal called Designing for the Sixth
Extinction explores the idea of putting synthetic living creatures into the
wild to save endangered species and clean up pollution. Ginsberg states, "The idea is that we could preserve or maintain a
state of nature using synthetic organisms that are designed to save other
species." Four new species have been proposed including a slug that
leaves a trail of alkali to neutralize acidic soil and a porcupine with sticky
rubber spines that would help disperse seeds of threatened plants. There is also an artificial puffball that
kills tree-damaging pathogens when it bursts; and a biofilm that grows on
leaves and absorbs pollutants and viruses, safely removing them when the leaves
fall in autumn. The creatures would be engineered to contain a genetic
"kill switch" that would prevent them from over-breeding and creating
new environmental problems.http://www.dezeen.com/2013/11/13/synthetic-creatures-could-save-nature-says-alexandra-daisy-ginsberg/
http://news.discovery.com/tech/biotechnology/synthetic-creatures-could-save-the-planet-131115.htm
Friday, November 23, 2012
Designed genetic circuit may force cancer cells to commit suicide
The researchers that took part in this project designed this genetic circuit by studying a type of newly discovered genetic material known as microRNA and choosing that to be their "target." MicroRNA, according to MIT News, is a snippet of RNA that helps regulate gene expression by selectively destroys messenger RNA. The reason why these researchers chose microRNA was because a large amount of particular types were found in cancer cells. According to the article, each different kind of cancer contained its own microRNA profile.
The microRNA profile of the cervical cancer the researchers chose to experiment with, HeLa, contained six microRNAs that were able to be identified. They were found in large quantities as well as unique to this particular cancer. They then created a synthetic gene that would code for a protein that would trigger apoptosis, hBax. This gene could be turned off by high levels of microRNA that are usually found low in HeLa as well as by low levels of microRNA that are usually found in large amounts in HeLa. However, this research is still fairly new and still being tested. According to these researchers, this process needs to be tested on living animals. While this system detects up to five characteristics in a cancer cell, they are still working on trying to get the system to identify more markers.
It is one step closer to finding another form of cancer treatment. Despite the fact it is still in its early stages and is not readily available to be used as a form of treatment just yet, it is still amazing that these researchers are able to design and program a gene to do this, while still keeping healthy cells unharmed. If this were to be a readily form of cancer treatment, would it be able to detect pre-cancerous cells rather than just cells that are already in their cancerous state? With further study and experimentation, I believe that they will be able to revise the design so that the circuit can detect cancerous cells earlier on and use this system for other kinds of cancer, not just this particular cervical cancer.
Subscribe to:
Comments (Atom)