:max_bytes(150000):strip_icc()/cortisol-371314fd2ef34947969e615ee1e75691.jpg)
Punnett's Square
Genetics news & views from students enrolled in BIOL 2110 at Stockton University.
Saturday, April 27, 2024
Genetic Risk Factors for Anxiety
:max_bytes(150000):strip_icc()/cortisol-371314fd2ef34947969e615ee1e75691.jpg)
Friday, April 26, 2024
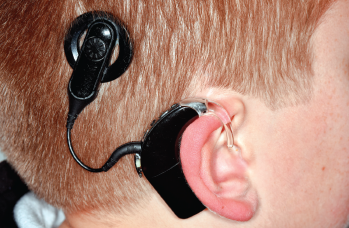
Gene Therapy Brings Hope to Children with Congenital Deafness
Epigenetic Factors in Drosophila are Capable of Causing Cancer Without Any Somatic Mutation.
Octopuses and Cephalopods Can Edit Their RNA
New research by Joshua Rosenthal and Eli Eisenberg identified that octopuses and other cephalopods adjust to environmental differences like temperatures by editing their RNA. Previous research has found that cells have the capacity to swap one member of the four letter genetic code, Adenosine, for a substitute molecule, Inosine. This protein altering is called A-to-I and it was found in octopuses. Researchers used the California two-spot octopus and acclimated them to their natural range of temperatures in the cold and warmer tanks. When examining their RNA they found increases at 13,285 sites in the cold tanks where the one letter change alters the protein. In the warmer tanks they found 550 sites. With the help of other collaborators at the University of Michigan and Texas Tech University, they were able to identify proteins that were altered when the processes of RNA editing occurred. One of the proteins was kinesin-1, which changes the rate at which this molecule travels. This then alters the responsiveness of a protein called synaptotagmin that allows for communication between neurons.
Mutation Helps Those at Higher Risk of Dementia avoid Alzheimer's
Redrawing The Tree of Life
In the article called “Huge Genetic Study Redraws The Tree of Life for Flowering Plants”, research has found that the origins of flowering and fruit-bearing plants are most likely all interconnected to each other. Molecular probes were designed to detect 353 genes that are also found in angiosperms. Charles Darwin considered the abundance of angiosperms an “abominable mystery” because it was not quite clear how they emerged. However, new technology has found over 9,500 species of flowering plants to have come from a great angiosperm bloom 140 million years ago. This gives scientists a better understanding of the tree of life.
In my opinion, having a better understanding of the tree of life can be very environmentally beneficial. This can help us improve conservation efforts, especially for plant species that might be going extinct. We can also find ways to sustainably farm fruit-bearing plants when we understand their relation to the tree of life and how they flourish. We can also use these plants for pharmaceutical research and medical practices as well.
Scientists find chemical that stops locust cannibalism
Locusts are generally solitary creatures. However, when their population density skyrockets due to heavy rainfall and good breeding conditions, followed immediately by an increase in food scarcity, a hormonal rush causes them to swarm and become aggressive. Fear of cannibalism drives the swam forward in one direction, as stopping will cause them to be eaten by the individual behind them.
In an experiment, scientists discovered there are specific scents produced exclusively during this hormonal phase, one of which repels other locusts, preventing them from being cannibalized. They believe this finding could be used in future methods of locust control, steering them away from farmland and crops.
This article really caught my eye. We have all heard of the horror caused by swarms of locusts, but I don't think we really think about why the locusts exhibit this behavior. The idea of them fearing being cannibalized as what drives the swarm is both horrifying and somewhat funny, albeit in a weird way. I do think it's incredible, however, that we now potentially have a way to prevent swarms of locusts from destroying food that could be feeding starving communities. What an interesting discovery!
Article: Scientists find chemical that stops locust cannibalism
Additional Information: A chemical defense deters cannibalism in migratory locusts
Thursday, April 25, 2024
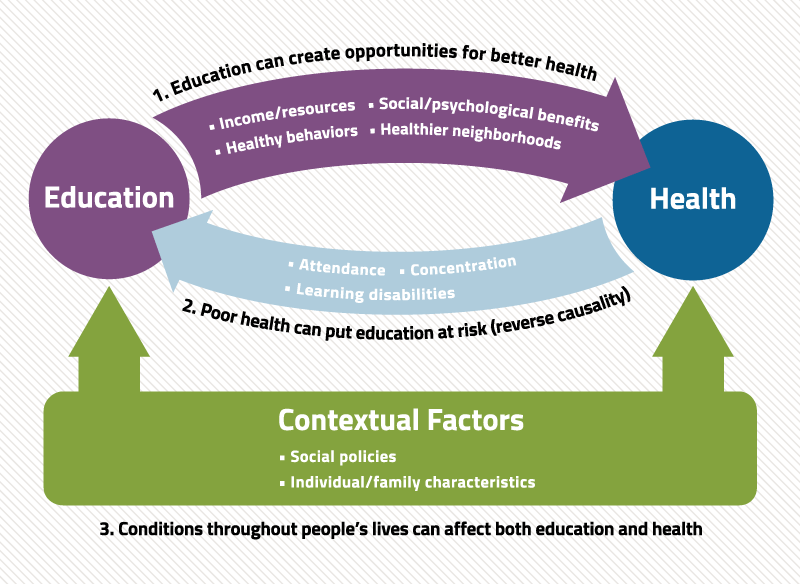
Epigenetics and Education
This article on genetics and learning evaluates the connection between genetics and educational attainment. Educational attainment is the highest level of education completed by an individual. The author finds that the mental processes needed for learning are often genetic in nature. He directly challenges previous held idea about education and development stating:
"Children are not a simple tabula rasa, as once thought, since they are conditioned by their genome to an extent. The environment...is also a significant factor which can allow them to make the most of their capabilities..."
Genetics play a big role in who we become in every aspect of our lives. This, however, does not mean we are bound to it. Oddly enough, our environment is also not capable of completely masking or rewriting the traits we enter the world with. As far as education goes, genetics are the tools we begin our journey with, but our environment has the ability to sharpen or dull them. This author advocates for further research into the neurological functions behind learning and environmental impact on education Together these studies can be used to create flexable and personalized learning plans that improve student outcomes. On the opposite end of the spectrum, I believe it can be used to identify outdated and potentially harmful teaching practices.
Overall, I believe this information is very important to America and the world as a whole. Many people are unhappy with their schools systems, each having unique problems that can interfere with a students learning abilities. For example:
Do standardized tests contribute more to high achievement or competition and surface level engagement?
How has Americas history with school shooters changed the average students attitude towards school environments and education?
Gen Z is notoriously called lazy and uneducated, but is that the truth or do they no longer seek educational attainment in favor of more basic needs?
Hopefully, we will have the answers to some of our education questions in the near future and the ability to make the needed changes for everyones best interest!